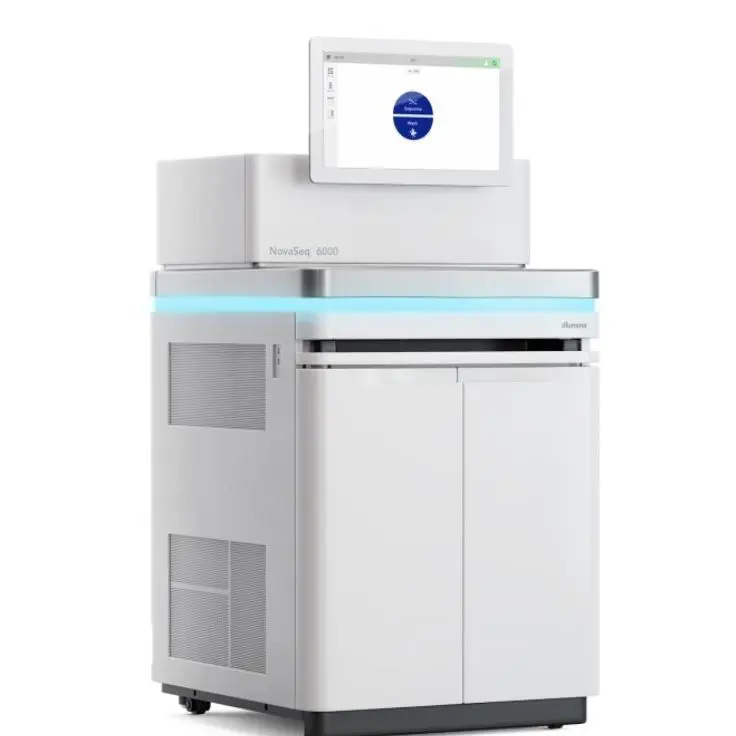

Illumina NovaSeq 6000 massive parallel sequencer combines two-color chemistry with patterned flow cells to enable high-throughput and high-accuracy sequencing, delivering ≥ 85% of sequenced bases with a quality score of Q30 or higher. It allows a modular and scalable approach to sequencing, with the availability of several sequencing formats, up to 20B clusters and 6 TB of data per run (dual flowcell) and up to 2×250 read lengths. It is suited for projects of different sizes, providing both the necessary high-throughput for large-scale sequencing projects, as well as the flexibility to accommodate small and medium-size projects at relatively moderate costs. At Candiolo Genomics we currently have one NovaSeq 6000 instrument.
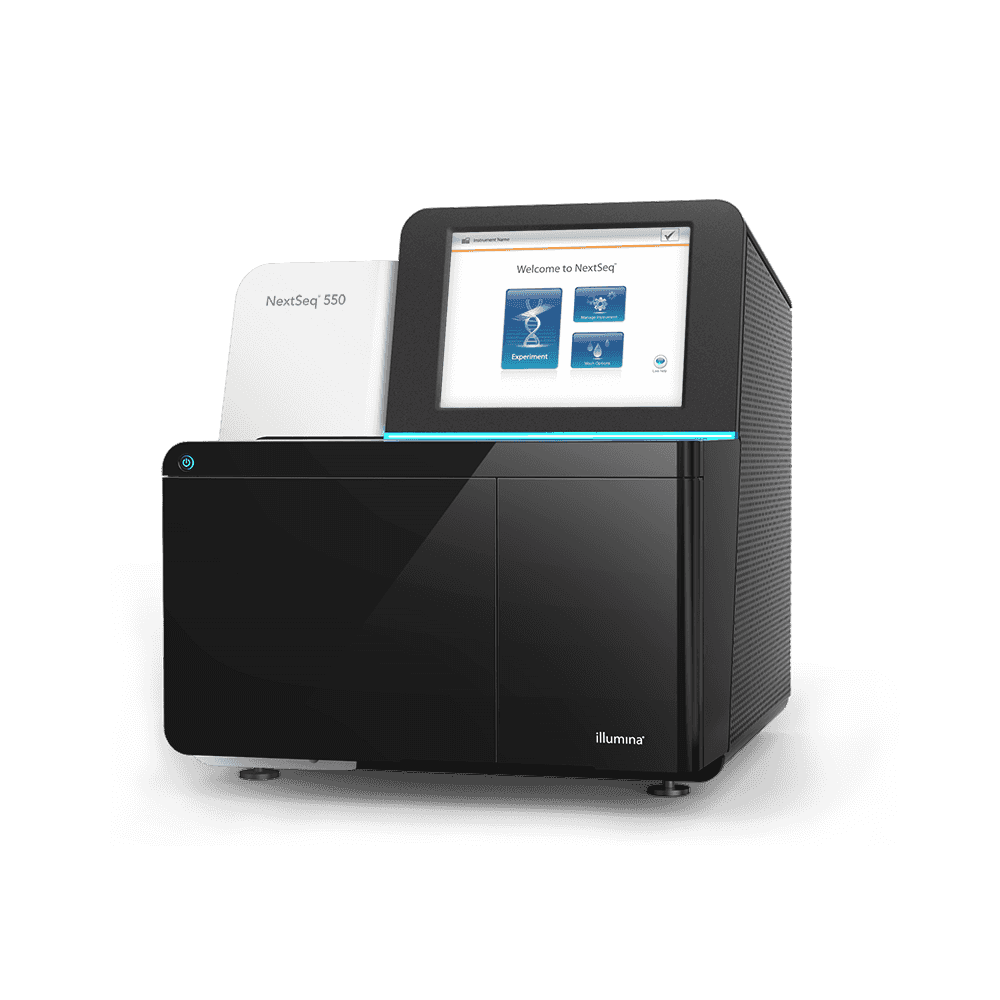

Illumina NextSeq500 Sequencing System is suited for medium to low-throughput projects, with an output range up to 400M clusters and 120 Gb per run, and up to 2×150 read lengths. At Candiolo Genomics we currently have one NextSeq500 instrument.
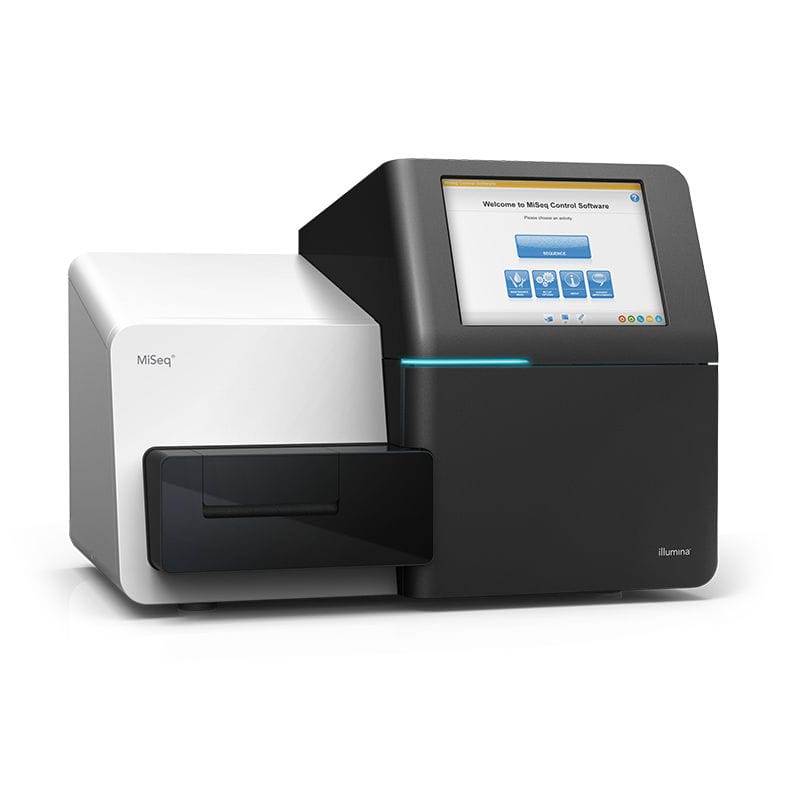

Illumina MiSeq Sequencing System can accommodate a variety of applications, including targeted gene sequencing, small genome sequencing, amplicon sequencing, 16S metagenomics analysis, with an output range up to 25M clusters and 15 Gb per run, and up to 2×300 read lengths. At Candiolo Genomics we currently have two MiSeq instruments.

Oxford Nanopore Technologies GridION and PromethION-24 Platforms support sequencing of long fragments of DNA and RNA using a proprietary array of nanopores embedded in an electro-resistant membrane. The systems provide real-time sequence information and full operator control of the desired output. Both instruments allow PCR-free sequencing of native DNA or RNA molecules with easy identification of DNA/RNA modifications and can also enable the identification of DNA/RNA modifications including 5mC, 5hmC, and m6A in DNA and m6A in RNA. Their modular asset with multiple independent flow cells allows for multiple experiments to be conducted in parallel. At Candiolo Genomics we currently have one GridION and one PromethION-24 instruments.

The 10X Genomics Chromium X quickly and efficiently partitions and barcode hundreds to over a million fresh, frozen, or fixed cells in a single run. Its advanced microfluidics provides efficiency, reproducibility, and performance in generating single cell partitions in just minutes, with each partition containing an identifying barcode for downstream analysis.
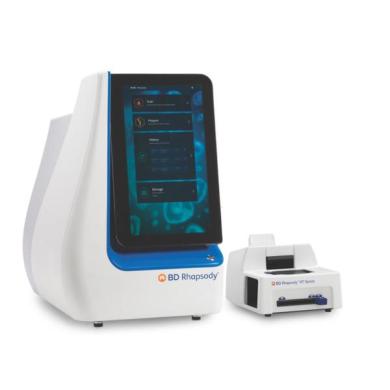
The BD Rhapsody Single-Cell Analysis System allows flexible sample processing and cell capture from hundreds to hundreds of thousands of single-cells using a microwell-based cartridge technology and multitier barcoding system.
Multiple samples can be processed in a single run when utilizing BD multiplexing antibodies. The captured cellular information is utilized to generate various types of libraries for next-generation sequencing applications providing accelerated time to insight.

The 10X Genomics Visium CytAssist is a powerful platform for Spatial Transcriptomic discovery, enabling the transfer of transcriptomic probes from standard glass slides to Visium slides, enabling spatial profiling insights to be gained from even more samples, such as FFPE blocks, presectioned FFPE, Fresh Frozen, or Fixed Frozen tissues on glass slides. Compatible with hematoxylin and eosin (H&E)- or immunofluorescence (IF)-stained tissue sections, CytAssist allows pre-sectioned tissues to be used for the Visium workflow.

QC and Automation
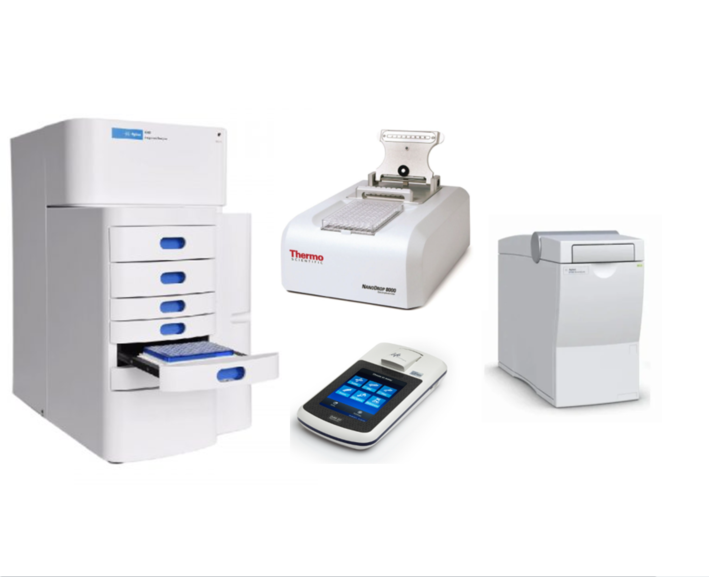
The quality and quantity of DNA and RNA prior to library preparation, as well as the intermediate and final library, are carefully checked by:
– The Nanodrop spectrometer for purity (A260/280 A 260/230 absorbance ratios)
– The Qubit Fluorometer for fluorescent accurate quantitation
– The Agilent 2100 Bioanalyzer and Agilent Fragment Analyzer for assessing DNA and RNA integrity prior to library preparation and to check the final library size.
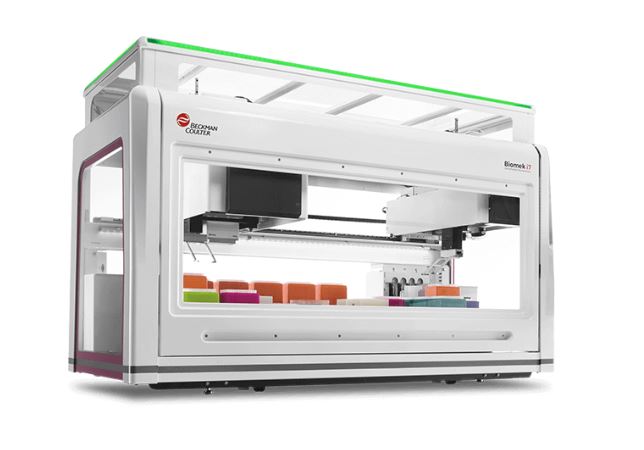

Automated library preparation on Beckman Coulter’s Biomek i7 is used to increase consistency and scalability, and ensure reproducible results